Sara Walker and Her Crew Publish the Most Interesting Biology Paper of 2022 (So Far, Anyway)
We’ve just ended the first quarter of the year. It’s a long way to New Year’s Eve 2022. But this new open access paper from senior author Sara Walker (Arizona State) and her collaborators will be hard to top, in the “Wow, that is so interesting!” category. (The first author of this paper is Dylan Gagler, so we’ll refer to it as “Gagler et al. 2022” below.)
1. Back in the day, the best evidence for a single Tree of Life, rooted in the Last Universal Common Ancestor (LUCA), was the apparent biochemical and molecular universality of Earth life.
Leading neo-Darwinian Theodosius Dobzhansky expressed this point eloquently in his famous 1973 essay, “Nothing in biology makes sense except in the light of evolution”:
The unity of life is no less remarkable than its diversity…Not only is the DNA-RNA genetic code universal, but so is the method of translation of the sequences of the “letters” in DNA-RNA into sequences of amino acids in proteins. The same 20 amino acids compose countless different proteins in all, or at least in most, organisms. Different amino acids are coded by one to six nucleotide triplets in DNA and RNA. And the biochemical universals extend beyond the genetic code and its translation into proteins: striking uniformities prevail in the cellular metabolism of the most diverse living beings. Adenosine triphosphate, biotin, riboflavin, hemes, pyridoxin, vitamins K and B12, and folic acid implement metabolic processes everywhere. What do these biochemical or biologic universals mean? They suggest that life arose from inanimate matter only once and that all organisms, no matter now diverse, in other respects, conserve the basic features of the primordial life.[Emphasis added.]
For Dobzhansky, as for all neo-Darwinians (by definition), the apparent molecular universality of life on Earth confirmed Darwin’s prediction that all organisms “have descended from some one primordial form, into which life was first breathed” (1859, 494) — an entity now known as the Last Universal Common Ancestor, or LUCA. So strong is the pull of this apparent universality, rooted in LUCA, that any other historical geometry seems unimaginable.
The “Laws of Life”
Theoretician Sara Walker and her team of collaborators, however, are looking for an account of what they call (in Gagler et al. 2022) the “laws of life” that would apply “to all possible biochemistries” — including organisms found elsewhere in the universe, if any exist. To that end, they wanted to know if the molecular universality explained under neo-Darwinian theory as material descent from LUCA (a) really exists, and (b) if not, what patterns do exist, and how might those be explained without presupposing a single common ancestor.
And a single common ancestor, LUCA? That’s what they didn’t find.
2. Count up the different enzyme functions — and then map that number within the total functional space.
Many thousands of different enzyme functional classes, necessary for the living state, have been described and catalogued in the Enzyme Commission Classification, according to their designated EC numbers. These designators have four digits, corresponding to progressively more specific functional classes. For instance, consider the enzyme tyrosine-tRNA ligase. Its EC number, 6.1.1.1, indicates a nested set of classes: EC 6 comprises the ligases (bond-forming enzymes); EC 6.1, those ligases forming carbon-oxygen bonds; 6.1.1, ligases forming aminoacyl-tRNA and related compounds; finally, 6.1.1.1, the specific ligases forming tyrosine tRNA. (See Figure 1.)
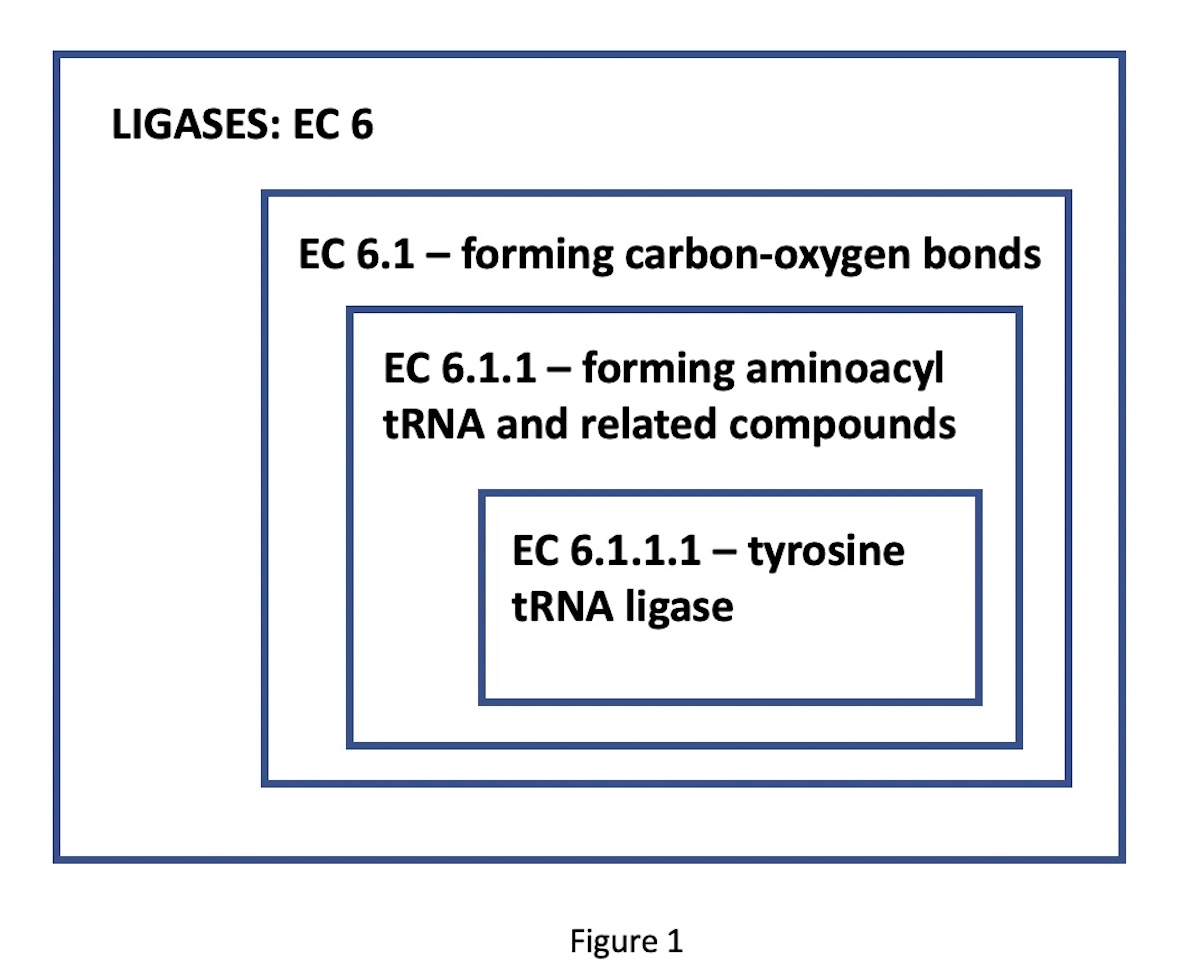
The Main Takeaway from This Pattern?
Being a ligase — namely, an enzyme that forms bonds using ATP — entails belonging to a functional group, but not a group with material identity among its members. A rough parallel to a natural language such as English may be helpful. Suppose you wanted to express the idea of “darkness” or “darkened” (i.e., the relative absence of light). English supplies a wide range of synonyms for “darkened,” such as:
- murky
- shaded
- shadowed
- dimmed
- obscured
The same would be the case — the existence of a set of synonyms, i.e., words with the same general meaning, but not the same sequence identity — for any other idea. The concept of something being “blocked,” for instance, takes the synonyms:
- jammed
- occluded
- prevented
- obstructed
- hindered
While these words convey (approximately) the same meaning, and hence fall into the same semantic functional classes, they are not the same character strings. Their locations in an English dictionary, ordered by alphabet sequence, may be hundreds of pages apart. Moreover, as studied by the discipline of comparative philology, the historical roots of a word such as “hindered” will diverge radically from its functional synonyms, such as “blocked.” These two words, although semantically largely synonymous, enter English from originally divergent or unrelated antecedents — a character string gap still reflected by their very different spellings.
A strikingly similar pattern obtains with the critical (essential) components of all organisms. Gagler et al. 2022 looked at the abundances of enzyme functions across the three major domains of life (Bacteria, Archaea, Eukarya), as well as in metagenomes (environmentally sampled DNA). What they found was remarkable — a finding (see below) which may be easier for non-biological readers to understand via another analogy.
3. A segue into computer architectures — then back to enzymes.
The basic architecture of laptop computers includes components present in any such machine, defined by their functional roles:
- Central processing unit (CPU) — the primary logic operator
- Memory — storage of coded information
- Power supply — electrons (energy) needed for anything at all to be computed
And so on. (Although exploring this point in detail would take us far afield, it is worth noting that in 1936, when Alan Turing defined a universal computational machine, he did so with no idea about the arrival, decades down the road, of silicon-based integrated circuits, miniaturized transistors, motherboards, solid-state memory devices, or any of the rest of the material parts of computers now so familiar to us. Rather, his parts were functionally, not materially defined, as abstractions occupying the various roles those parts would play in the computational process — whatever their material instantiation would later turn out to be.) Now suppose we examined 100,000 laptops, randomly sampled from around the United States, to see what type of CPU — meaning which material part (e.g., built by which manufacturer) — each machine used as its primary logic operator.
A range of outcomes is possible (see Figures 2A and 2B). For instance, if we plot CPUs from different manufacturers on the y axis, against the total number of laptop parts inspected on the x axis, it might be the case that the distribution of differently manufactured (i.e., materially distinct) CPUs would scale linearly with laptops inspected (Figure 2A). In other words, as our sample of inspected laptop parts grows, the number of different CPUs discovered would trend upwards correspondingly.
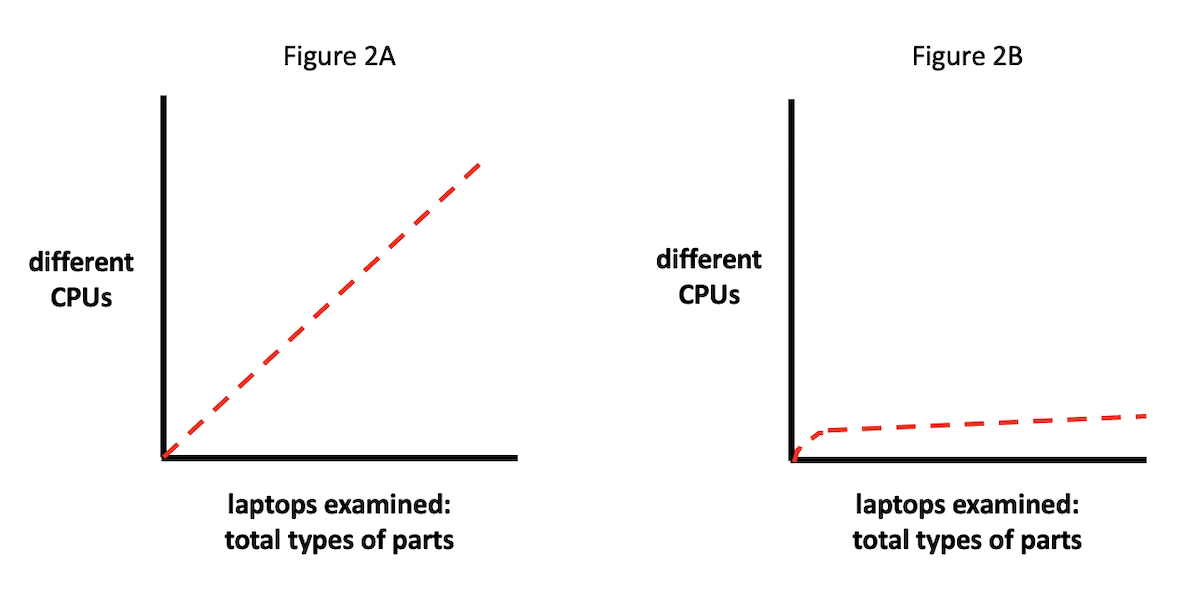
Or — and this fits, of course, with the actual situation we find (see Figure 2B) — most of the laptops would contain CPUs manufactured either by Intel or AMD. In this case, we would plot a line whose slope would change much more slowly, staying largely flat, in fact, after the CPUs from Intel and AMD were tallied.
The Core Rationale of Their Approach
Now consider Figure 3 (below), from the Gagler et al. 2022 paper. This shows the core rationale of their approach: tally the EC-classified enzyme “parts” within each of the major domains, and from metagenomes, and then plot that tally against the total EC numbers.
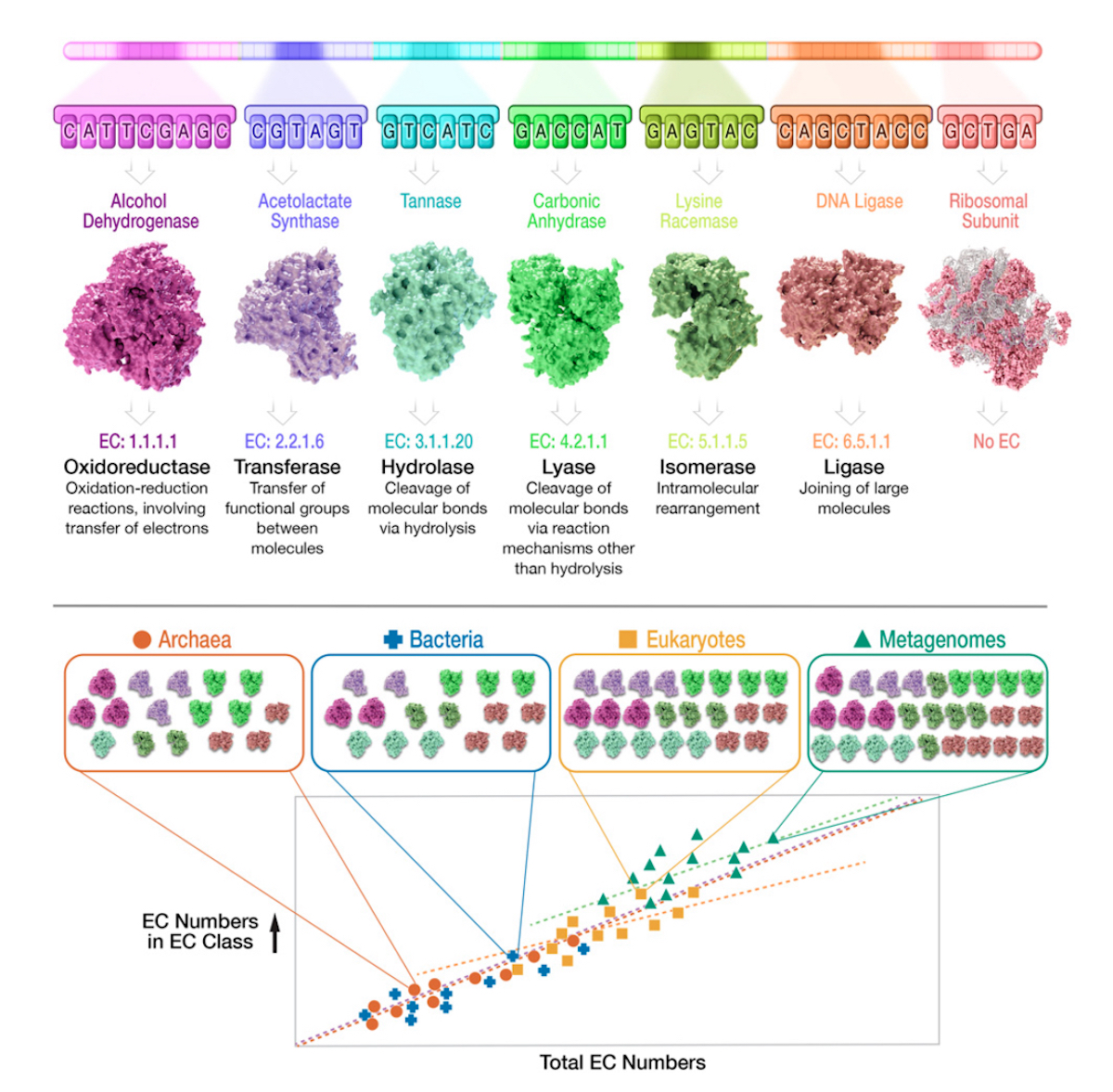
Figure 3 also shows their main finding. As the enzyme reaction space grows (on the horizontal axis — total EC numbers), so do the number of unique functions (on the vertical axis — EC numbers in each EC class).
The lesson that Gagler et al. 2022 draw from this discovery? The pattern is NOT due to material descent from a single common ancestor, LUCA. Indeed, under the heading, “Universality in Scaling of Enzyme Function Is Not Explained by Universally Shared Components,” they explain that material descent from LUCA would entail shared “microscale features,” meaning “specific molecules and reactions used by all life,” or “shared component chemistry across systems.” If we use the CPU / laptop analogy, this microscale commonality would be equivalent to finding CPUs from the same manufacturer, with the same internal logic circuits, in every laptop we examine.
But what Gagler et al. 2022 found was a macroscale pattern, “which does not directly correlate with a high degree of microscale universality,” and “cannot be explained directly by the universality of the underlying component functions.” In an accompanying news story, project co-author Chris Kempes, of the Santa Fe Institute, described their main finding in terms of functional synonyms: macroscale functions are required, but not the identical lower-level components:
“Here we find that you get these scaling relationships without needing to conserve exact membership. You need a certain number of transferases, but not particular transferases,” says SFI Professor Chris Kempes, a co-author on the paper. “There are a lot [of] ‘synonyms,’ and those synonyms scale in systematic ways.”
As Gagler et al. frame the point in the paper itself (emphasis added):
A critical question is whether the universality classes identified herein are a product of the shared ancestry of life. A limitation of the traditional view of biochemical universality is that universality can only be explained in terms of evolutionary contingency and shared history, which challenges our ability to generalize beyond the singular ancestry of life as we know it. …Instead, we showed here that universality classes are not directly correlated with component universality, which is indicative that it emerges as a macroscopic regularity in the large-scale statistics of catalytic functional diversity. Furthermore, EC universality cannot simply be explained due to phylogenetic relatedness since the range of total enzyme functions spans two orders of magnitude, evidencing a wide coverage of genomic diversity.
Sounds Like Intelligent Design
It is interesting to note that this paper was edited (for the PNAS) by Eugene Koonin of the National Center for Biotechnology Information. For many years, Koonin has argued in his own work that the putative “universality due to ancestry” premise of neo-Darwinian theory no longer holds, due in large measure to what he and others have termed “non-orthologous gene displacement” (NOGD). NOGD is a pervasive pattern of the use of functional synonyms — enzyme functions being carried out by different molecular actors — in different species. In 2016, Koonin wrote:
As the genome database grows, it is becoming clear that NOGD reaches across most of the functional systems and pathways such that there are very few functions that are truly “monomorphic”, i.e. represented by genes from the same orthologous lineage in all organisms that are endowed with these functions. Accordingly, the universal core of life has shrunk almost to the point of vanishing…there is no universal genetic core of life, owing to the (near) ubiquity of NOGD.
Universal functional requirements, but without the identity of material components — sounds like design.
No comments:
Post a Comment